A new coronavirus outbreak emerged on the 31st of December 2019 in Wuhan, China, causing commotion among the medical community and the rest of the world. This new species of coronavirus has been termed 2019-nCoV and has caused a considerable number of cases of infection and deaths in China and, to a growing degree, beyond China, becoming a worldwide public health emergency. 2019-nCoV has high homology to other pathogenic coronaviruses, such as those originating from bat-related zoonosis (SARS-CoV), which caused approximately 646 deaths in China at the start of the decade. The mortality rate for 2019-nCoV is not as high (approximately 2–3%), but its rapid propagation has resulted in the activation of protocols to stop its spread. This pathogen has the potential to become a pandemic. It is therefore vital to follow the personal care recommendations issued by the World Health Organization.
Un nuevo brote de coronavirus surgió el pasado 31 de diciembre de 2019 en Wuhan, China, causando conmoción entre la comunidad médica y el resto del mundo. Esta nueva especie de coronavirus fue denominada como 2019-nCoV, causante de un gran número de casos y fallecimientos en China y en cantidad creciente fuera de ella, convirtiéndose en una emergencia de salud pública a nivel mundial. 2019-nCoV es un virus con alta homología con otros coronavirus patogénicos, como los originados por zoonosis con murciélagos (SARS-CoV) causantes de aproximadamente 646 muertes en China a principios de la década. Su tasa de mortalidad no es tan elevada (aproximadamente del 2–3%), pero su rápida propagación ha propiciado la activación de protocolos para detener su diseminación. Este patógeno tiene el potencial para convertirse en pandemia, por lo que es vital seguir las recomendaciones de cuidado personal dictadas por la Organización Mundial de la Salud.
In December 2019, a series of cases of pneumonia caused by a novel coronavirus were identified in the city of Wuhan, China. This novel coronavirus has various names: 2019-nCoV (according to the World Health Organization) and SARS-CoV-2 (according to the International Committee on Taxonomy of Viruses). The disease that this virus causes has been called 2019-nCoV.1 On the 7th of January 2020, the novel coronavirus was officially announced by Chinese authorities as the causal agent of these infections.1
The coronaviruses are enveloped in nonsegmented positive-sense RNA that belong to the Coronaviridae family and Nidovirales order and are widely distributed among humans and mammals, causing numerous conditions that range from the “common” influenza to death.2
As of the 13th of February 2020, several cases have been recorded. According to the World Health Organization (WHO), there have been 46,997 cases reported worldwide, 46,550 of which (99.04%) have been confirmed in China. Of these, 1368 (2.93%) resulted in death, thereby categorizing this disease as a worldwide public health emergency.3,4
The following link provides an real-time updated map on the worldwide situation of cases of coronavirus, with incidence rates and locations: https://gisanddata.maps.arcgis.com/apps/opsdashboard/index.html -/bda7594740fd40299423467b48e9ecf6.
Structural characteristics of coronavirusesCoronaviruses can be differentiated into 4 genera: alpha, beta, delta and gamma. To date, the alpha and beta types have been determined to infect humans,5 causing diseases ranging from the common cold to more severe conditions, such as Middle East respiratory syndrome (MERS-CoV) and severe acute respiratory syndrome (SARS-CoV), which caused thousands of deaths in 2002.6,7 Four of the human coronaviruses (HCoV 229E, NL63, OC43 and HKU1) are endemic worldwide and represent 10–30% of upper respiratory tract infections in adults.5
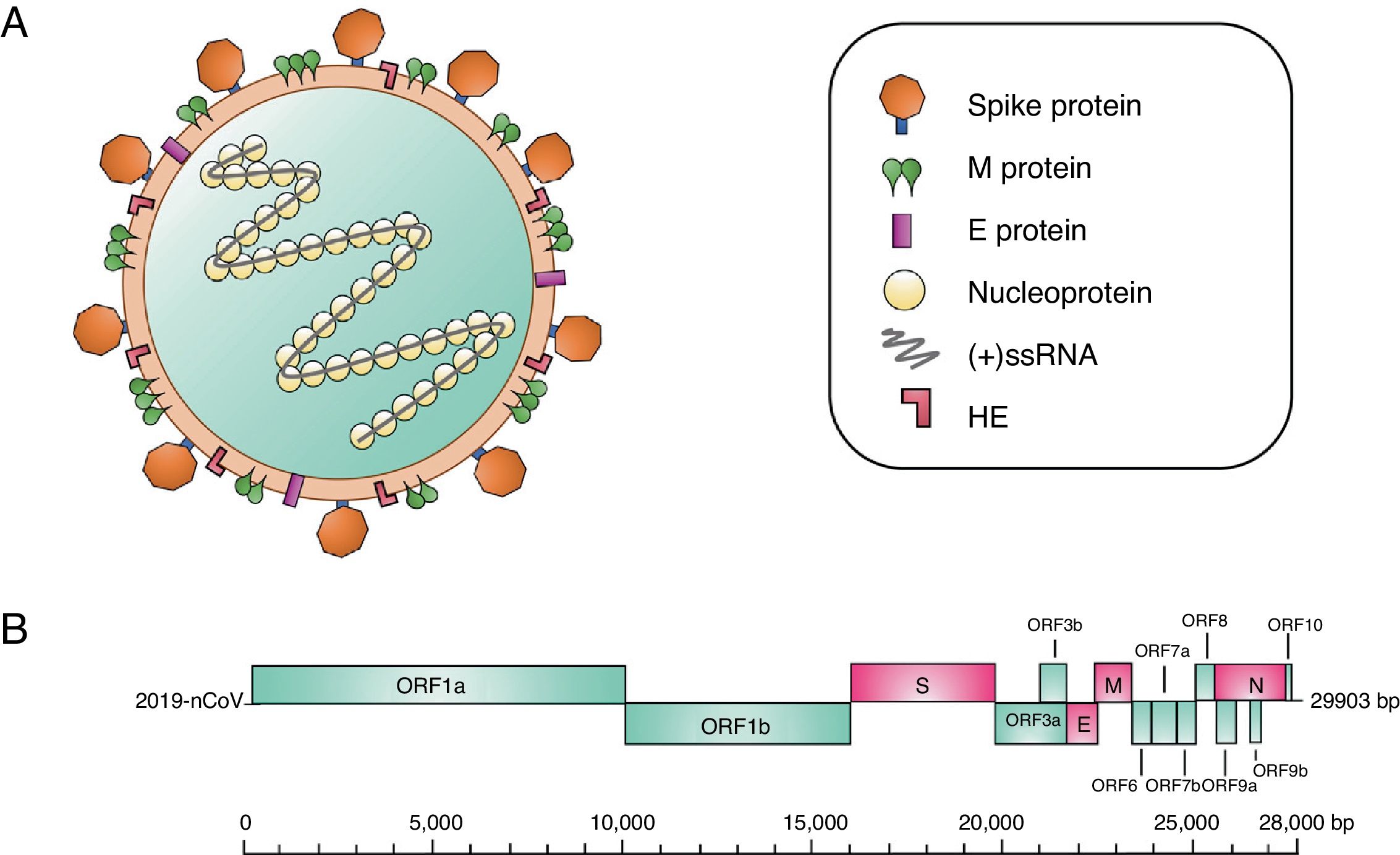
Structurally, coronaviruses are spherical or pleomorphic, with diameters of 80–120nm. Various electronic microscope analyses have identified the surface of the virion, discovering that they are structures organized by projections that in turn consist of viral spike (S) glycoprotein trimers.8 Additionally, other short projections composed of hemagglutinin-esterase (HE) protein dimers have been identified, which have been observed in some beta-coronaviruses.9 For its part, the viral envelope is reinforced by the membrane (M) glycoprotein (the most abundant on the virion's surface), which is found embedded in the membrane by 3 transmembrane domains.10 Another structural component of the virion is the envelope (E) protein, a small protein that is highly hydrophobic and is found to a small proportion than the other proteins.11 The viral proteins of the coronavirus are embedded in a lipid membrane that is created by the infected cells. Internally, the virus particle consists of an additional protein known as the nucleoprotein (N), which binds to the virus RNA in a helical structure similar to a beaded string, thereby protecting the RNA from degradation (Fig. 1A).8
(A) Coronavirus particle. This type of virus contains positive-polarity single-strand RNA [(+)ssRNA] genetic material, measuring 27–32kilobases. The virus consists of a nucleocapsid, which in turn is composed of (+)ssRNA and the nucleoprotein. This structure is covered by a lipid bilayer. Other structural proteins of the coronavirus are found here, such as the spike protein, which covers this virus particle, as well as hemagglutinin-esterase (HE) dimers, the highly hydrophobic envelope (E) protein and the membrane (M) protein, the most abundant protein on the virion's surface. (B) Organization of the genes in the 2019-nCoV genome. The structural genes are shown in pink, and the nonstructural genes are shown in blue.
The genome of coronaviruses is positive-polarity, single-strand nonsegmented RNA (+ssRNA), which measures 27 to 32kilobases. The genomic RNA presents certain changes such as polyadenylations in the 3’ terminal region; in contrast, the 5’ terminal region contains a cap structure.12 Within this RNA, there are numerous open reading frames (6–11 ORFs). The first ORF encodes approximately 16 nonstructural proteins, while the remaining ORFs encode accessory and nonstructural proteins (Fig. 1B).13
The genome sequence analyses for 2019-nCoV were fairly similar to those of SARS-CoV; however, there are certain differences such as the lack of the region encoding for protein 8a in 2019-nCoV, which could therefore have resulted in its lower pathogenesis when compared with SARS-CoV.14
OriginOne unknown that is still being investigated is the virus’ zoonotic origin. Due to the close similarity between 2019-nCoV and bat coronaviruses, it is likely that bats are the main reservoir for the virus. Various studies were conducted when this new class of coronavirus appeared and discovered that 2019-nCoV is 96% identical to a bat coronavirus at the genome level. The same study revealed that this virus belongs to the SARS-CoV species.15 Thus, it was speculated that SARS-CoV was transmitted to humans from exotic animals in markets in the outbreak 18 years ago,15 while MERS-CoV was transmitted from camels to humans.16
The reports documented that many of the first identified patients share the common factor of contact with a seafood and animal market; however, other patients had no contact with these locations at any time, which shows the limited person-to-person infection after identifying groups of cases among families, as well as transmission from patients to health workers.17 Moreover, a recently published study estimated that 95% of the cases of 2019-nCoV infections in Wuhan showed symptoms before the 12th of January 2020,18,19 a fact that, combined with the virus’ incubation period, suggests a high possibility of the disease's travel-related propagation.20,21
TransmissionGenerally, coronaviruses replicate primarily in the epithelial cells of the lower respiratory tract and, to a lesser extent, in the cells of the upper respiratory tract. Transmission therefore mainly occurs from patients with recognized disease and not from patients with mild and nonspecific signs; in other words, it is believed that propagation occurs only after the patient shows signs of the disease in the lower respiratory tract.22,23 However, patients with 2019-nCoV infection detected in a severe or fatal state have a greater probability of transmitting this virus, given that they expel a greater quantity of infectious particles than patients who have the mild or asymptomatic form of the infection. Identifying and quarantining these patients in health centers where outbreaks have occurred, along with the implementation of appropriate infection control and constant reports on cases from various countries, have been effective in reducing the transmission and containing outbreaks of the disease.24
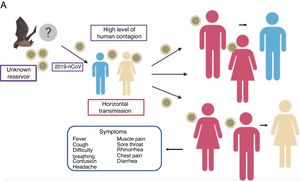
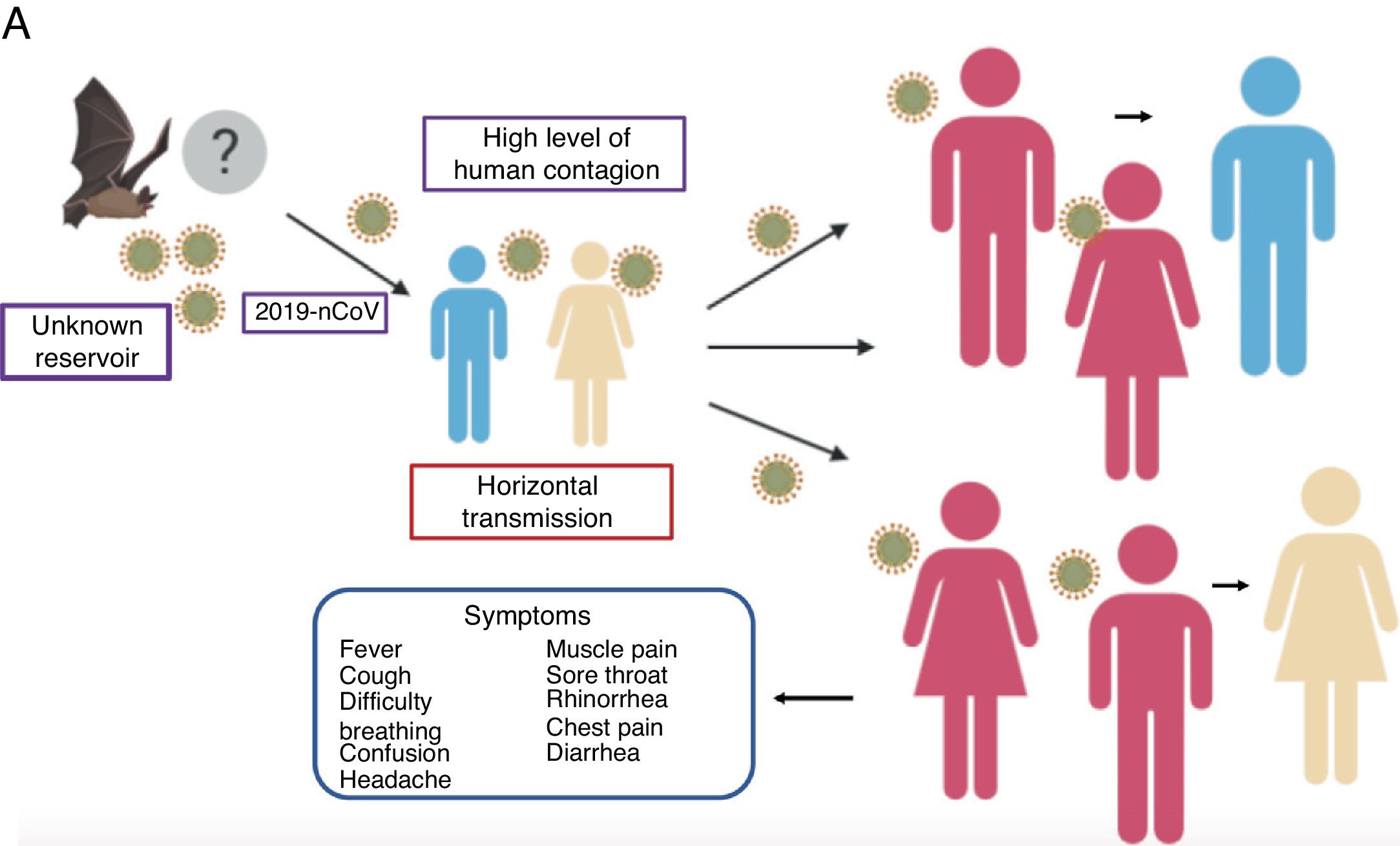
The mean number of new cases that a case of coronavirus generates over the course of its infectious period (R0) varies between 2.24 (95% CI 1.96–2.55) and 3.58 (95% CI 2.89–4.39),21 i.e., one individual can infect approximately 2 to 4 people, which means that the infection can propagate rapidly and widely among the population (Fig. 2).
Propagation of 2019-nCoV. There are various hypotheses regarding the virus’ animal-to-human transmission, the strongest of which is the bat origin. Human-to-human transmission has been reported to occur through the respiratory tract, thereby facilitating infection between populations. The symptoms are very common, with fever, cough, muscle pain and respiratory problems, among others.
Who are most susceptible to the infection? The novel coronavirus can infect people of all ages, although the elderly and those with pre-existing medical conditions (such as asthma, diabetes and heart disease) seem to be most vulnerable to falling seriously ill with the virus. Thus, the reported mortality rate for individuals older than 70 years is >8%. According to reports, the patients who died of this infection had a mean age of 56 years, and most had other diseases (heart disease, stroke, diabetes, etc.), which could have made them more vulnerable to the virus. The Chinese Center for Disease Control and Prevention reported that 1 to 2 men were infected for every woman. It has been speculated that women's low susceptibility to the viral infections could be due to protection from their “extra” X chromosome.25 Children constitute a peculiar population with an immune system different from that of adults. Virus transmission through their mothers with suspected or confirmed infection therefore occurs easily.26 However, a lower severity and extremely low mortality has been observed.
Clinical presentationThe diagnosis of the disease has been reached based on virus-induced pneumonia based on the clinical symptoms observed in the patients (similar to that of other respiratory viruses), to their history of exposure to other people with the virus, and the history of visits to areas affected.7,27 Moreover, a detailed study of the first 99 patients treated in the Wuhan Jinyintan Hospital, conducted between the 1st and 20th of January 2020, showed that of the 99 patients with 2019-nCoV pneumonia, 49% had a history of exposure to the Huanan seafood market and that 51% had chronic diseases. These patients’ mean age was 55.5 years, 67 were men and 32 were women, and the incubation period for the virus was estimated at 7–14 days.27 The analyzed patients had clinical manifestations of fever (83%), cough (82%), difficulty breathing (31%), muscle pain (11%), confusion (9), headache (8%), sore throat (5%), runny nose (4%), chest pain (2%), diarrhea (2%), nausea and vomiting (1%). According to imaging tests, 75% of these patients showed bilateral pneumonia, 14% showed significant mottling and ground-glass opacity, and 1% had pneumothorax. Similarly, 17% developed acute respiratory distress syndrome, of whom 11% worsened in a short period and died due to multiple organ failure.23,28 Moreover, the leukocyte counts were below the normal range in 9 of the patients and above the normal range in 24; 38% of them had above normal neutrophil counts. The lymphocyte and hemoglobin counts were below the normal range for many of the patients, and the platelet counts were below the normal range in 12% of the patients and above normal in 4%. Forty-three of these patients had varying degrees of impaired hepatic function, with alanine aminotransferase (ALT) or aspartate aminotransferase (AST) levels above the normal range. One patient experienced severe hepatic function impairment.28,29
Classification of patients according to their clinical symptomsConsidering the patients’ clinical symptoms, when should a 2019-nCoV case be labeled as suspected, probable or confirmed? According to the WHO, a suspected case involves a patient with severe acute respiratory infection (fever, cough, requiring hospitalization) and with no other etiology that completely explains the clinical presentation, as well as a history of travel or residence in China during the 14 days before symptom onset. Alternatively, a suspected case can involve a patient with an acute respiratory disease and at least one of the following factors during the 14 days prior to the onset of the symptoms: (a) contact with a confirmed or probable case of 2019-nCoV infection or (b) working or attending a medical care center that was treating patients with confirmed or probable 2019-nCoV infection and patients with acute respiratory disease. A probable case involves a patient whose 2019-nCoV tests are not conclusive or are positive using an assay for coronavirus but no laboratory evidence of other respiratory pathogens. Finally, confirmed cases involve patients with positive laboratory results for 2019-nCoV infection, regardless of the signs and clinical symptoms.30
Clinical tests for the diagnosisAmong the main priorities for facilitating public health interventions in patients is the laboratory diagnosis. In the case of an acute respiratory infection, the reverse transcription-polymerase chain reaction is commonly employed to identify the viruses that cause the respiratory secretions.30,31
The test for detecting the viral envelope's gene sequence has been effectively implemented by 35 laboratories; however, the diagnostic algorithm employs other sequences of the viral genome to confirm positivity for 2019-nCoV by detecting gene sequences of the viral RNA polymerase and nucleoprotein.32
Other samples to collect include…
- (a)
RNA extraction from clinical samples using the MagNA Pure 96 system.
- (b)
Respiratory material (nasopharyngeal and oropharyngeal swabs for outpatients and sputum [if presented] and/or endotracheal aspirate or bronchoalveolar lavage for patients with severe respiratory disease).33
- (c)
Serum for serological tests, acute samples and convalescent samples (in addition to the respiratory material).33
The objectives of the diagnostic tests are to detect the common causes of early pneumonia, to support disease control activities and to work with reference laboratories that can perform pancoronavirus detection and direct sequencing.31
Prevention measuresThe standard recommendations issued by the WHO for preventing the propagation of the infection include washing hands on a regular basis, especially after contact with sick individuals or their environment, covering the mouth and nose when coughing and sneezing, cooking meat and eggs thoroughly, avoiding close contact with any individual who presents symptoms of respiratory diseases (coughing and sneezing), avoiding traveling to affected cities and areas and avoiding close contact with live or dead farm animals or wild animals. Travelers with symptoms of acute respiratory infection should practice coughing etiquette (maintaining distance, covering the mouth when coughing and sneezing into clothing or disposable tissues and thoroughly washing hands).7,34,35
Those individuals who have had contact with patients categorized as having probable and/or confirmed 2019-nCoV infection should be monitored for 14 days from the last unprotected contact they had with infected individuals and should limit their travel to locations beyond their residence to avoid possible propagation.
One of the most widely used preventive measures is face masks, but can they stop the propagation? Surgical face masks for the public are not 100% effective protection against airborne viruses or bacteria, because the masks do not have an adequate air filter and leave the eyes exposed. Although face masks can help reduce the risk of contracting the virus through sneezing or coughing from others, the optimal solution is respirators with specialized air filters, because they are designed specifically to protect individuals against potentially dangerous airborne particles. These are known as filtering face piece (FFP) face masks, of the which there are 3 types:
- •
FFP1 – filters approximately 78% of air particles, thereby protecting against nontoxic and nonfibrogenic residues of dust and aerosols and preventing the inhalation of bothersome residues and smells.
- •
FFP2 – filters approximately 92% of air particles, protecting against nontoxic residues and fibrogenic elements, preventing the inhalation of toxic fluids of dust, aerosols and fumes.
- •
FFP3 – filters 98% of air particles, protecting against poisons and toxins of dust, smoke and aerosols, as well as bacteria, viruses and fungal spores.
For isolation procedures and where infectious aerosols (e.g., tuberculosis, measles, chickenpox, SARS) might be generated, the WHO recommends using a ventilator with a filtration efficiency of at least 95% for 0.3μm diameter particles, which is equivalent to an N95 facemask, according to the US National Institute for Occupational Safety and Health (NIOSH) regulations. Given that the US regulations differ from the European, this level of protection for the general population is between FFP2 and FFP3.
In general, the use of masks is recommended only if there is exposure to patients with respiratory diseases, such as in hospitals and medical offices. The guidelines for mask use should be strictly followed when exposing individuals to infected patients, because poor handling and touching the eyes with the hands and with the edge of the mask increase the risks.
Who should be notified if there are confirmed cases or an infection is suspected?Any individual who has been in contact with patients and who meets the clinical characteristics of the disease becomes a suspected case and should be confirmed through laboratory tests.30 The WHO requests that national authorities report confirmed cases of novel coronavirus infection within 24h of identifying them and provide the patients’ minimum basic data set.30 For Mexico, cases of patients with signs and symptoms of the disease can be reported to the Epidemiological and Health Intelligence Unit (Unidad de Inteligencia Epidemiológica y Sanitaria, UIES), 24h a day, 7 days a week, at (+52) 8000044800. In Spain, cases should be reported in Madrid to the Ministry of Health (Ministerio de Sanidad) at (+34) 915961000 or to the National Institute of Healthcare Management (Instituto Nacional de Gestión Sanitaria), at (+34) 913380000.
TreatmentMeanwhile, numerous research laboratories are currently seeking a treatment that can eliminate the MERS-CoV infection, either with the drugs that are already part of the pharmaceutical industry and are being employed for other diseases or by searching for new more specific alternatives for the virus, given that there is currently no specific vaccine or treatment, although there are several specific vaccines and treatments in the development phase.
Despite the short time, it has been predicted that a vaccine will be ready for animal testing in approximately a month and that in 3 months it could be ready for humans. For now, the treatment is only supportive and depends on the patient's clinical condition.36
Broad-spectrum antivirals, such as remdesivir, lopinavir/ritonavir and interferon beta, have shown promise against MERS-CoV in animal models, and their activity against the present 2019-nCoV is being evaluated.36 Scientists at the Vaccine Research Center of the National Institute of Allergy and Infectious Diseases of Washington DC have employed approaches that use nucleic acid-based vaccine platforms to counter the infection.37
ConclusionThe WHO is collaborating with specialists in public health and animal health, clinicians and scientists from around the world to meet and exchange scientific data to better understand the virus and the disease it causes. An example of these data is the identification of new secretion pathways in infected patients. Similarly, the WHO has focused on determining the priorities in the response to outbreaks, the therapeutic strategies for treatment and the approaches for clinical care. There are analyses that suggest that certain SARS-CoV-specific monoclonal antibodies can be effective in neutralizing 2019-nCoV.
Undoubtedly, it is everyone's task to help contain the outbreaks that have emerged in so short of a time. It is therefore advisable that the general population follow all recommendations for preventing the infection. Although the number of cases of infection outside of China are relatively low compared to the numbers within the country, every country should be prepared for any situation related to this virus.
Please cite this article as: Palacios Cruz M, Santos E, Velázquez Cervantes MA, León Juárez M. COVID-19, una emergencia de salud pública mundial. Rev Clin Esp. 2021;221:55–61.



![(A) Coronavirus particle. This type of virus contains positive-polarity single-strand RNA [(+)ssRNA] genetic material, measuring 27–32kilobases. The virus consists of a nucleocapsid, which in turn is composed of (+)ssRNA and the nucleoprotein. This structure is covered by a lipid bilayer. Other structural proteins of the coronavirus are found here, such as the spike protein, which covers this virus particle, as well as hemagglutinin-esterase (HE) dimers, the highly hydrophobic envelope (E) protein and the membrane (M) protein, the most abundant protein on the virion's surface. (B) Organization of the genes in the 2019-nCoV genome. The structural genes are shown in pink, and the nonstructural genes are shown in blue. (A) Coronavirus particle. This type of virus contains positive-polarity single-strand RNA [(+)ssRNA] genetic material, measuring 27–32kilobases. The virus consists of a nucleocapsid, which in turn is composed of (+)ssRNA and the nucleoprotein. This structure is covered by a lipid bilayer. Other structural proteins of the coronavirus are found here, such as the spike protein, which covers this virus particle, as well as hemagglutinin-esterase (HE) dimers, the highly hydrophobic envelope (E) protein and the membrane (M) protein, the most abundant protein on the virion's surface. (B) Organization of the genes in the 2019-nCoV genome. The structural genes are shown in pink, and the nonstructural genes are shown in blue.](https://static.elsevier.es/multimedia/22548874/0000022100000001/v1_202101120711/S2254887420300333/v1_202101120711/en/main.assets/thumbnail/gr1.jpeg?xkr=veFvTaG4yx3us0l+EPzH7Q==)